Globally, breast cancer causes around 670k deaths each year, making it one of the leading causes of cancer mortality among women. While governments and research institutions are taking steps to reduce these deaths, there is a gap in AI application. Neural networks have proven effective for medical predictions and can suggest treatments, but current research on neural networks lacks focus on medical applications to improve human lives.
Models like hugging-science have been created to bridge this gap, but many are not well documented, tested, or are just hobby projects. This is why we introduce breast-cancer-detector—a model trained carefully for optimal performance and great generalization. It was tested on an external dataset where it achieved a score of 96%.
Get the Model: breast-cancer-detector on Hugging Face
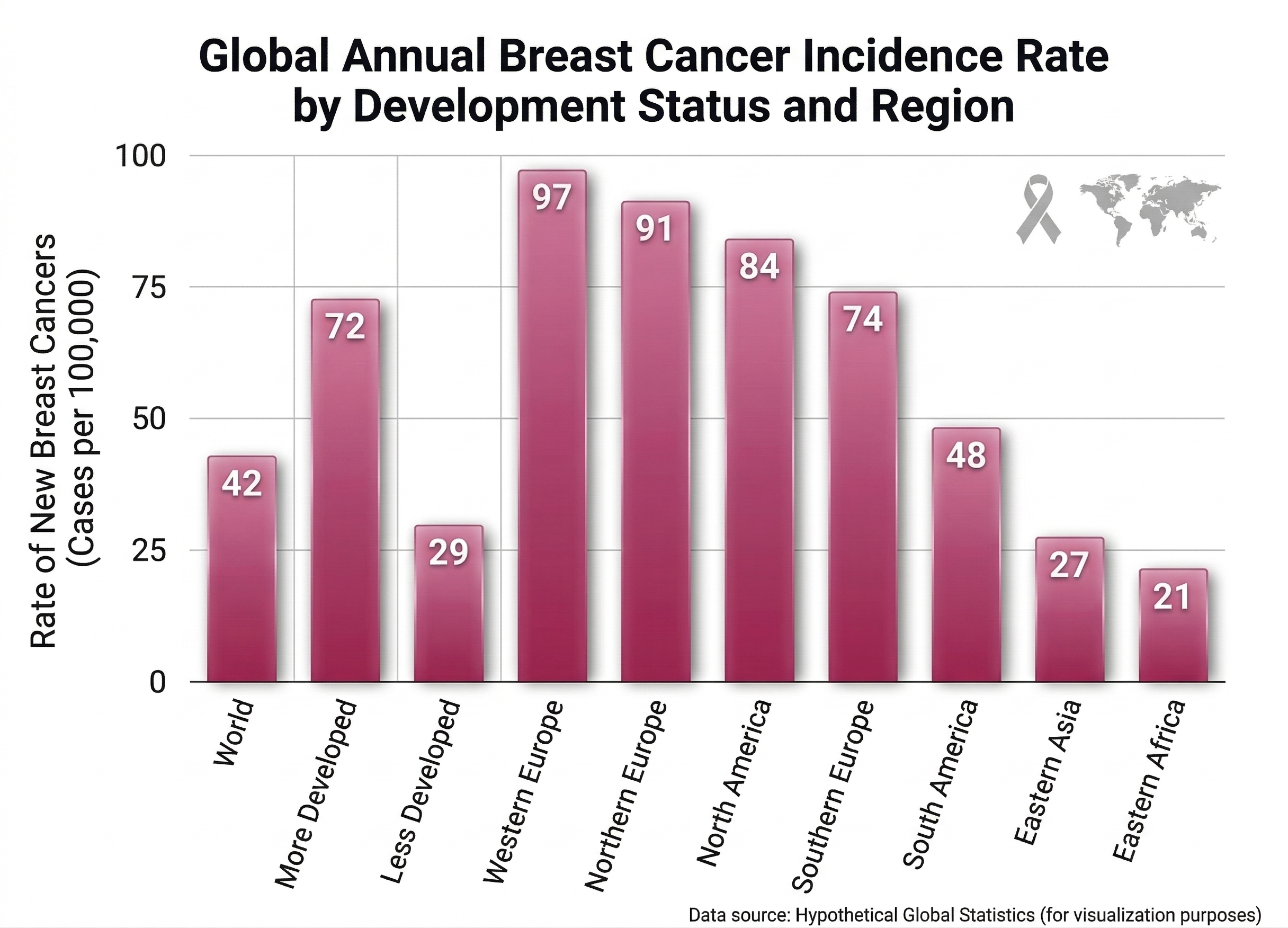
Statistics
About Model
This is a ViT (Vision Transformer) based model with around 86M parameters and a patch size of 16×16 (ref). Image size is set to 224px. You can find the base model here. The base model was just a feature extractor, and we converted it into a classifier by replacing the final pooling layer with a classification layer with 3 weights and 3 biases with a dimension of 768. This is the only change made to the architecture; everything else is the magic of data augmentation and training approach.
The three labels added were:
- 0: benign — Non-cancerous findings
- 1: malignant — Suspicious or cancerous findings
- 2: normal — No visible abnormalities
Training
The model was trained for 12 epochs on Breast-Cancer-Ultrasound-Images-Dataset, which contains only 1,500 samples but still gave us an impressive model. The loss in the 12th epoch was only 0.027 with an accuracy of 94% on the training set. To avoid overfitting, we added an extra 20% augmented data with noise, blur, and JPEG-like compression. This led the model to converge around 10-11 epochs without saturating early. We increased weight decay to 10% to prevent memorizing training data. The learning rate scheduler used was cosine, which gradually decreases the LR during training to stabilize the model. Despite the low LR of 5e-5, the training was done on an NVIDIA L4 with a batch size of 64.
You can find some extra info about training on the HF model card
Benchmarks and Evaluation
This model already demonstrates strong state-of-the-art performance in detection, which is why we decided to test it on an external dataset to validate its generalization.
We tested the model on the breastcanc-ultrasound-class dataset which is an external dataset containing 647 breast ultrasound images with only two classes (benign and malignant). Since this dataset doesn’t have a “normal” class like ours, we just mapped our model’s predictions accordingly where “benign” stays benign and “malignant” stays malignant. The “normal” predictions we got were only 3 out of 647 (0.46%) which is pretty cool because it shows the model understands that when it’s seeing actual lesions, it’s not gonna classify them as normal.
Out of the 647 images we tested, 644 had meaningful predictions (excluding the “normal” ones) and the results were pretty solid. We achieved an accuracy of 96.12% on this external dataset which is even better than our internal validation accuracy of 94.46% — this tells us the model is actually generalizing well and not overfitting to the training data.
The precision for malignant cases was 94.26% which means when we predict cancer, we’re usually right about it. The recall or sensitivity was 93.81% which means we’re catching most of the actual malignant cases that are there. The F1-score came out to 94.03% which is a good balance between precision and recall.
For benign cases, we had even better performance with a precision of 97.01% and recall of 97.24%. This is the kind of performance we want because both false positives and false negatives are risky in medical imaging — false negatives could miss cancer and false positives could cause unnecessary procedures.
{
"classification_report": [
{
"class": "benign",
"precision": 0.9701,
"recall": 0.9724,
"f1_score": 0.9712,
"support": 434
},
{
"class": "malignant",
"precision": 0.9426,
"recall": 0.9381,
"f1_score": 0.9403,
"support": 210
},
{
"class": "accuracy",
"precision": null,
"recall": null,
"f1_score": 0.9612,
"support": 644
}
]
}
This external validation really shows that our approach of using data augmentation with noise, blur, and JPEG compression worked because the model learned robust features that transfer well to completely new data it’s never seen before.
How to Use
Using the model is pretty straightforward. We hosted it on Hugging Face so you can use it directly via the transformers library. Just load the model as an image classification pipeline and pass your ultrasound image:
from transformers import pipeline
classifier = pipeline("image-classification", model="Parveshiiii/breast-cancer-detector")
# Pass the path/url to your ultrasound image
result = classifier("path/to/your/ultrasound/image.png")
print(result)
The model will return predictions with confidence scores for all three classes. The output will look something like this:
[
{"label": "benign", "score": 0.95},
{"label": "malignant", "score": 0.04},
{"label": "normal", "score": 0.01}
]
Make sure you’re passing clean ultrasound images without any text overlays, annotations, or other artifacts — the model was trained on clean images so it performs best on those. You can also use it with .jpg or .jpeg files, not just PNG.
If you want to integrate it into a web app, you can use Gradio or Streamlit for quick prototyping.
One of the checkpoints from this project has been donated to Hugging Face Science to support open medical AI research. You can find it here.